 |
The Telomere is comprised of double strand DNA (dsDNA) that repeats every 6 bases that continues on into single strand DNA (ssDNA). Because a portion of the telomere is single stranded, it is important that the cells also protect it from repair systems that would wrongly act on it. To protect these ends - the telomere forms a "T-loop" structure where the ssDNA loops back and inter-twines with the dsDNA. The purpose of the experiments of this paper is to show evidence of a protein TRF2 (Telomere Repeat binding Factor protein) that may help to form these protective structures. In this report, Scientific AmeriKen will focus in on its contribution to that paper. 
TRF1 and TRF2 are two proteins that are known to bind to telomere DNA and protect them. Although the two proteins are very similar there is an extra region that exist in TRF2 that does not exist in TRF1. Additionally, it is known that TRF2 tends to be located near the ends of the telomere while TRF1 is found near the beginning. Finally, earlier experiments in the paper seem to indicate that TRF2 binds differently to test DNA comprised of different lengths of dsDNA/ssDNA. Based on these data, our hypothesis is that TRF2 may bind directly to the ds-ss DNA junction.

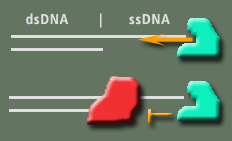 Certain proteins exist that simply digest DNA. A commonly used technique, called "DNA footprinting", takes advantage of the abilities of these proteins (called nucleases) to determine if and where another protein of interest might be binding to the DNA. The idea is if the protein of interest binds the DNA it will prevent the nuclease from digesting it (image on right). In this experiment Exonuclease I was used. This nuclease binds specifically to the ends of ssDNA and digest base by base inward. Our goal will be to digest DNA with Exo I by itself and in the presence of TRF2 to see if TRF2 can stop its activity. Certain proteins exist that simply digest DNA. A commonly used technique, called "DNA footprinting", takes advantage of the abilities of these proteins (called nucleases) to determine if and where another protein of interest might be binding to the DNA. The idea is if the protein of interest binds the DNA it will prevent the nuclease from digesting it (image on right). In this experiment Exonuclease I was used. This nuclease binds specifically to the ends of ssDNA and digest base by base inward. Our goal will be to digest DNA with Exo I by itself and in the presence of TRF2 to see if TRF2 can stop its activity.
To perform the experiment first DNA had to be labeled with radioactive Phosphorus (P32). This allows for the DNA to be visualized through electrophoresis. Next, two tubes are set up, one with DNA, the other with DNA and TRF2. After some time is given to allow TRF2 to bind the DNA - a sample is removed to record time=0. Next Exonuclease I is added and samples are taken at specific time points. These samples are then examined via electrophoresis to determine the effects of TRF.

Figure A is shown as a simple control to demonstrate that the DNA is in fact double stranded. Preparation of the DNA includes first taking the longer strand and making it radioactive. Then the shorter strand is added, and then heated to cause all DNA to come apart. They are slowly cooled to allow them to bind together again. The result of this "annealing" can be see through electrophoresis - in this case, the unannealed band appears lower than the annealed band - this implies the annealed band is of larger size - as would be expected since it has two DNA strands.
A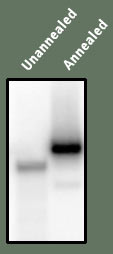 | | B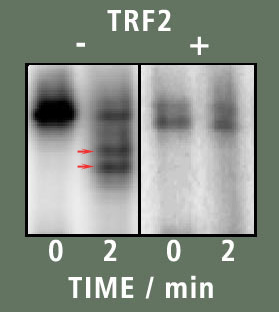 |
In figure B the result of the experiment is seen. The left box contained the sample that has no TRF2 in it. From it we can see that after 2 minutes, Exo I digested a considerable portion of the DNA. This is seen by the existence of bands below the starting band that are seen in the two minute lane (two red arrows). However when TRF2 is present, no such bands are present suggesting the DNA was not digested in the presence of TRF2.

From the data presented it appears as though TRF2 has some ability to protect the ssDNA region from the nuclease. Although one would have to ask - is TRF2 in fact targeting the nuclease and not the DNA? Could this be an effect of simply that there was a difference in the amount of DNA loaded in the two samples? What does the function or reason TRF2 binds to this location? Although more questions were created by this experiment - it does show Scientific AmeriKen's first data (although it was preliminary at this stage) to go on to make it into a paper! The new era begins!

The data from this experiment was preliminary - further experimentation was done with a different nuclease as Exonuclease I was stronger then what was needed. The efforts of Scientific AmeriKen can be seen in the paper, by Khan et. al., in figure 2A - although it is noted that the final data that is shown in the paper was completed by someone else. In total the paper used many techniques to demonstrate this interaction. The first was a mobility shift assay - showing that as TRF2 binding causes DNA to run slower in the electrophoresis process. Further mobility shifts demonstrate a large effect when you remove TRF2's unique extra portion to its ability to bind the DNA. Finally, using atomic force microscopy, TRF2 is actually visualized interacting with DNA. Atomic force microscopy is a process that uses an atomic sized needed to touch and scan out an image. In summary - these experiments provide evidence that TRF2 is a key protein in the protection of the telomeres. The abstract of this paper can be viewed by clicking here.
Scientific AmeriKen would like to thank Sheik Khan and Terace Fletcher for their help and training in this project. |
 |
|













 Certain proteins exist that simply digest DNA. A commonly used technique, called "DNA footprinting", takes advantage of the abilities of these proteins (called nucleases) to determine if and where another protein of interest might be binding to the DNA. The idea is if the protein of interest binds the DNA it will prevent the nuclease from digesting it (image on right). In this experiment Exonuclease I was used. This nuclease binds specifically to the ends of ssDNA and digest base by base inward. Our goal will be to digest DNA with Exo I by itself and in the presence of TRF2 to see if TRF2 can stop its activity.
Certain proteins exist that simply digest DNA. A commonly used technique, called "DNA footprinting", takes advantage of the abilities of these proteins (called nucleases) to determine if and where another protein of interest might be binding to the DNA. The idea is if the protein of interest binds the DNA it will prevent the nuclease from digesting it (image on right). In this experiment Exonuclease I was used. This nuclease binds specifically to the ends of ssDNA and digest base by base inward. Our goal will be to digest DNA with Exo I by itself and in the presence of TRF2 to see if TRF2 can stop its activity.
